
ECON 0150 | Economic Data Analysis
The economist’s data analysis skillset.
Part 4.3 | Model Assumptions
GLM (so far)
We’ve built three models. Can we trust them?
- Each model produces coefficients, standard errors, and p-values.
- Sometimes a linear model isn’t appropriate.
GLM Assumptions
Which model do you think offers better predictions?

GLM Assumptions
Which model do you think offers better predictions?

Model 1 has predictions close to the average data for all predictor values.
Model 2 will offer inaccurate predictions for large predictor variables!
GLM Assumptions
Our test results are only valid when the model assumptions are valid.
- Linearity: The relationship between X and Y is linear
- Homoskedasticity: Equal error variance across all values of X
- Independence: Observations are independent from each other
- Normality: Errors are normally distributed
GLM Assumptions: why check?
Assumption violations affect our inferences
If assumptions are violated:
- Coefficient estimates may be biased
- Standard errors may be wrong
- p-values may be misleading
- Predictions may be unreliable
We use residuals (\(\epsilon\)) and estimates (\(\hat{y}\)) to check if a model is ‘specified’.
Model Fit
We use residuals (\(\varepsilon\)) and estimates (\(\hat{y}\)) to check if a model is ‘specified’.

Residuals (\(\varepsilon\)): the error of each prediction; how far off the model is.
Estimates (\(\hat{y}\)): the model’s predicted value for each observation.
Exercise 4.3 | Happiness and GDP
Use a residual plot to visualize the error for the model.
Assumption 1: Linearity
The error term should be unrelated to the fitted value.
Q. Which model appears to violate the Linearity Assumption?
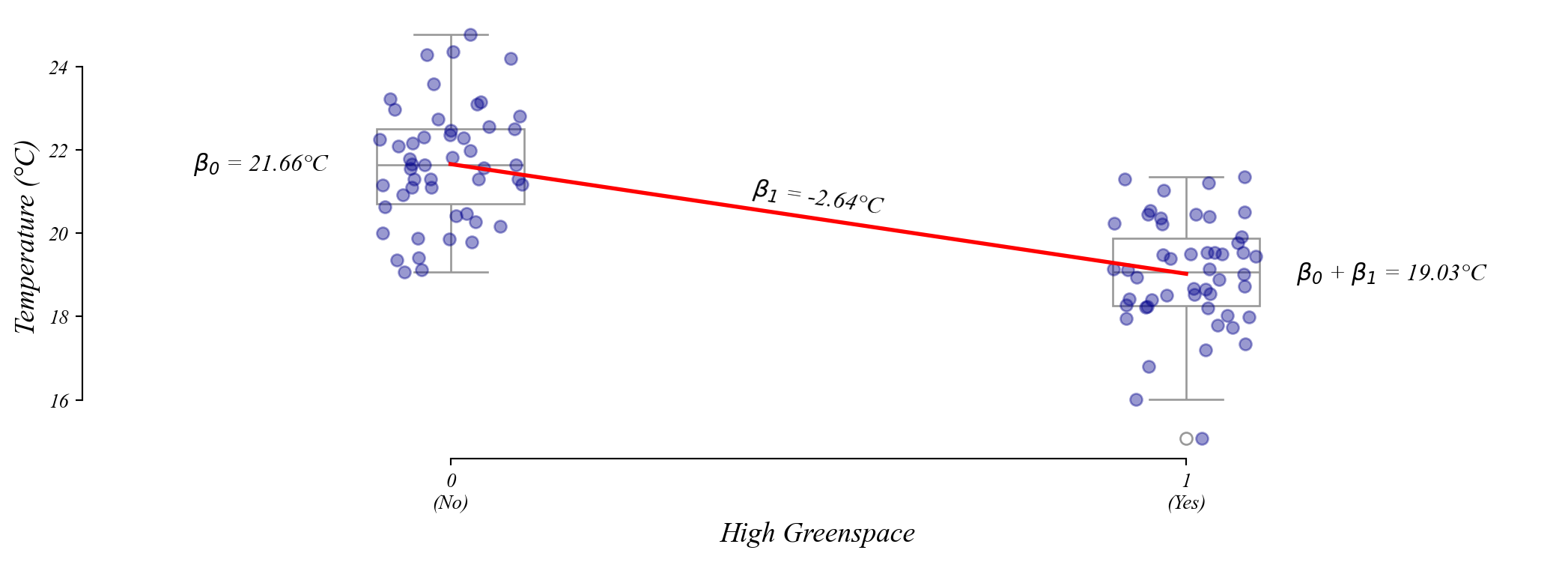
Assumption 1: Linearity
The error term should be unrelated to the fitted value.
Q. Which model appears to violate the Linearity Assumption?

Model 1 is equally wrong everywhere. Model 2 has larger errors at the extremes.
Assumption 1: Linearity
A non-linear relationship will produce non-linear residuals.
The linear model misses the true curvature leading to systematic errors.
Handling Non-Linear Relationships
Transform variables to become linear
Adding a square term or performing a log transformation can fix the problem.
instead of
\[\text{income} = \beta_0 + \beta_1 \text{age} + \varepsilon\]
we could use
\[\text{income} = \beta_0 + \beta_1 \text{age} + \beta_2 \text{age}^2 + \varepsilon\]
It’s also common to log transform either the \(x\) or \(y\) variable.
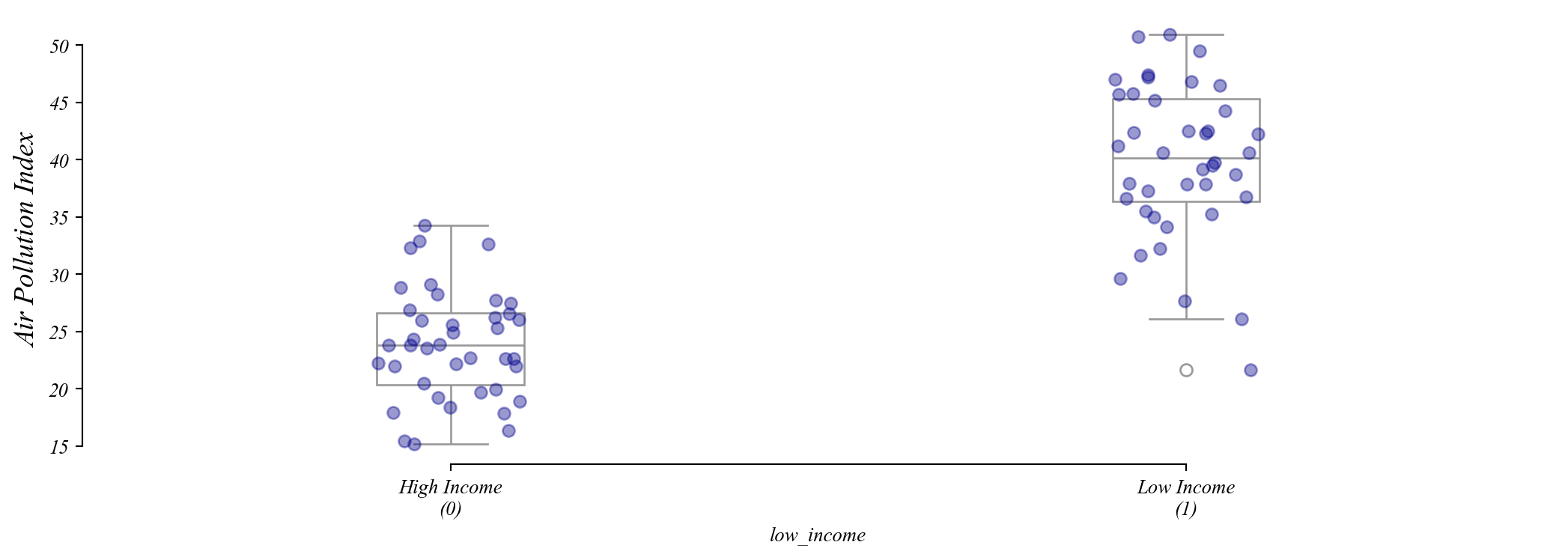
Assumption 2: Homoskedasticity
Residuals should be spread out the same everywhere.
Q. Which one of these figures shows homoskedasticity?
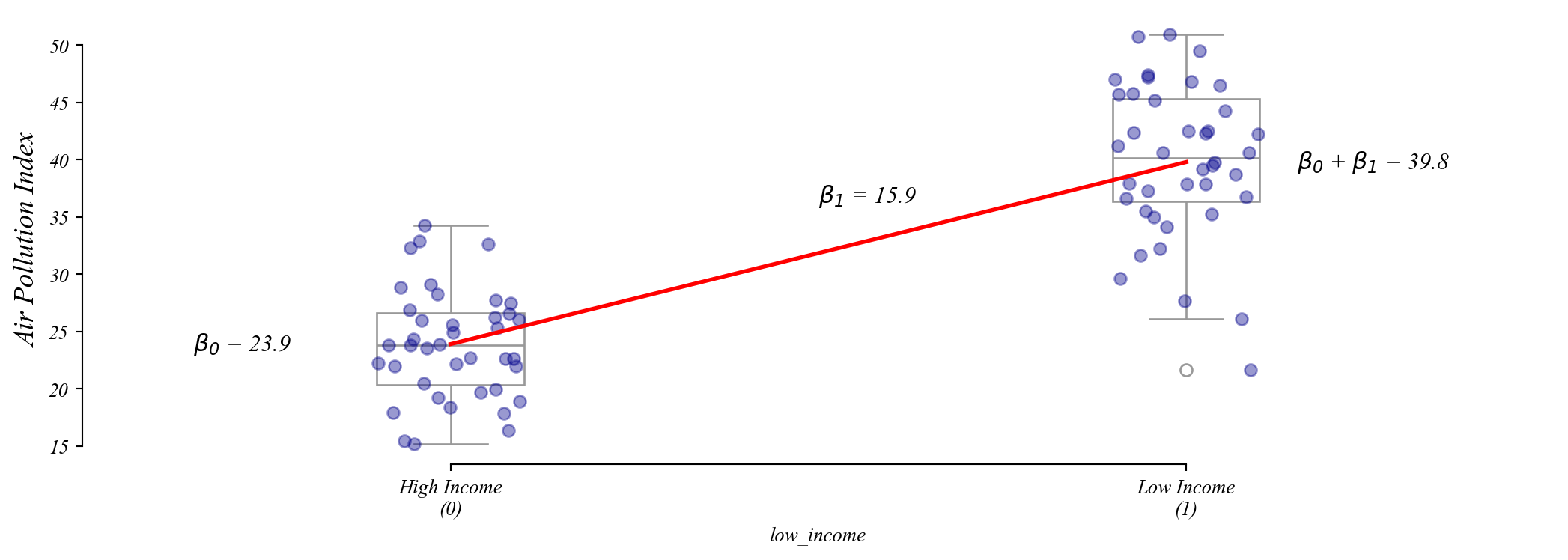
Assumption 2: Homoskedasticity
Residuals should be spread out the same everywhere.
Q. Which one of these figures shows homoskedasticity?
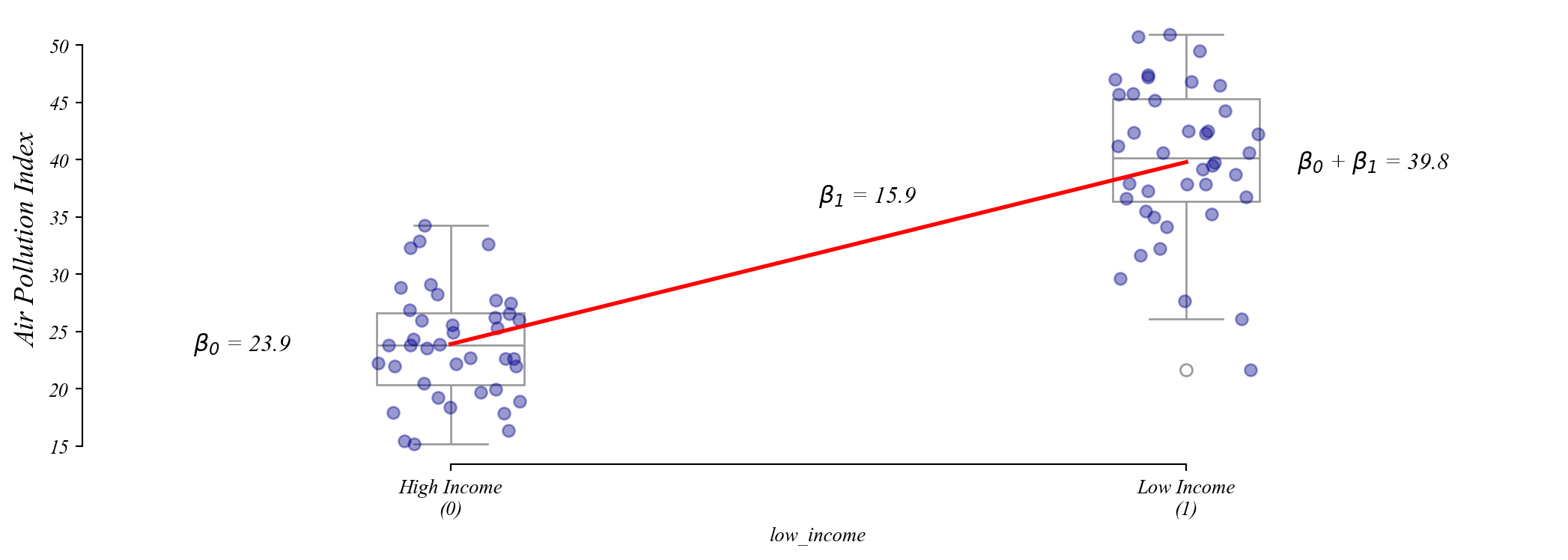
Model 1 is equally wrong everywhere. Model 2 has errors that grow with the fitted value.
Assumption 2: Homoskedasticity
The spread of residuals should not change across values of X.
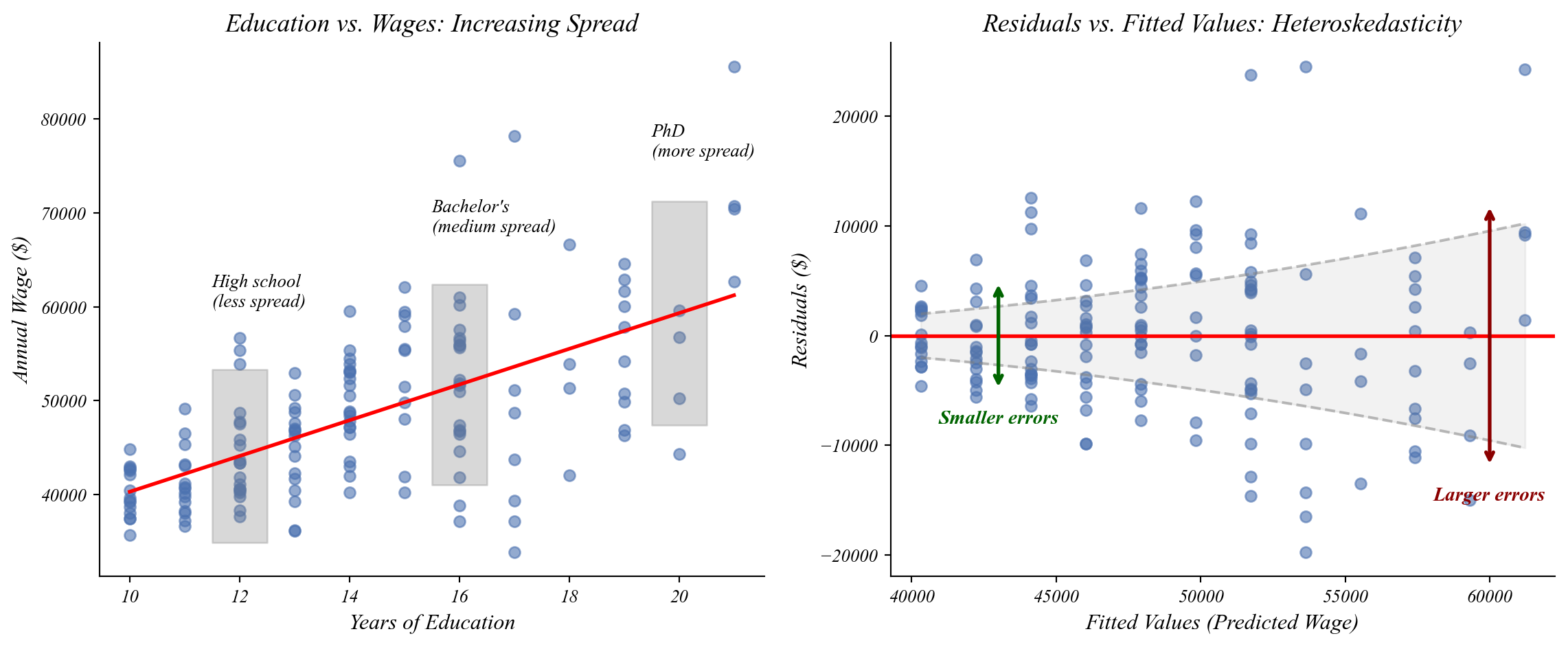
The spread of points increases as education increases!
Handling Heteroskedasticity
Robust standard errors give more accurate measures of uncertainty
Use Robust Standard Errors which adjust for the changing spread, giving more accurate p-values.
Assumption 3: Independence
Each error should be unrelated to previous errors.
Q. Which of these residual lag plots shows independence?
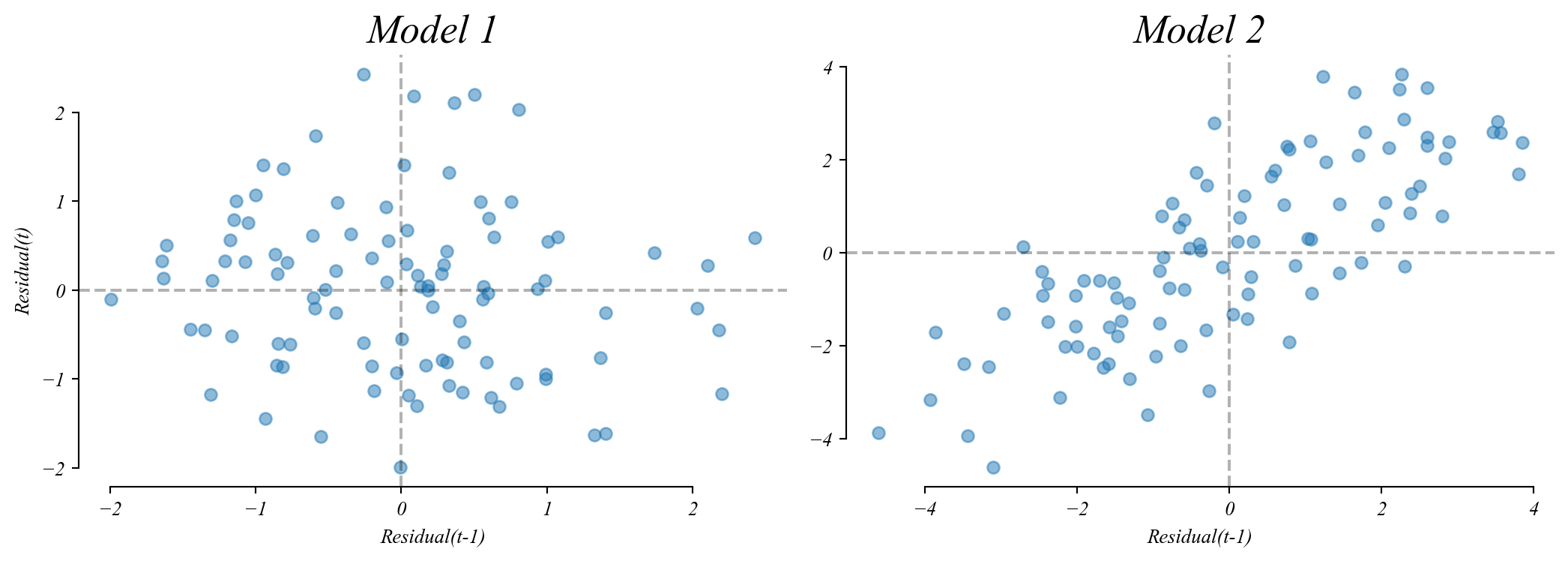
Assumption 3: Independence
Each error should be unrelated to previous errors.
Q. Which of these residual lag plots shows independence?
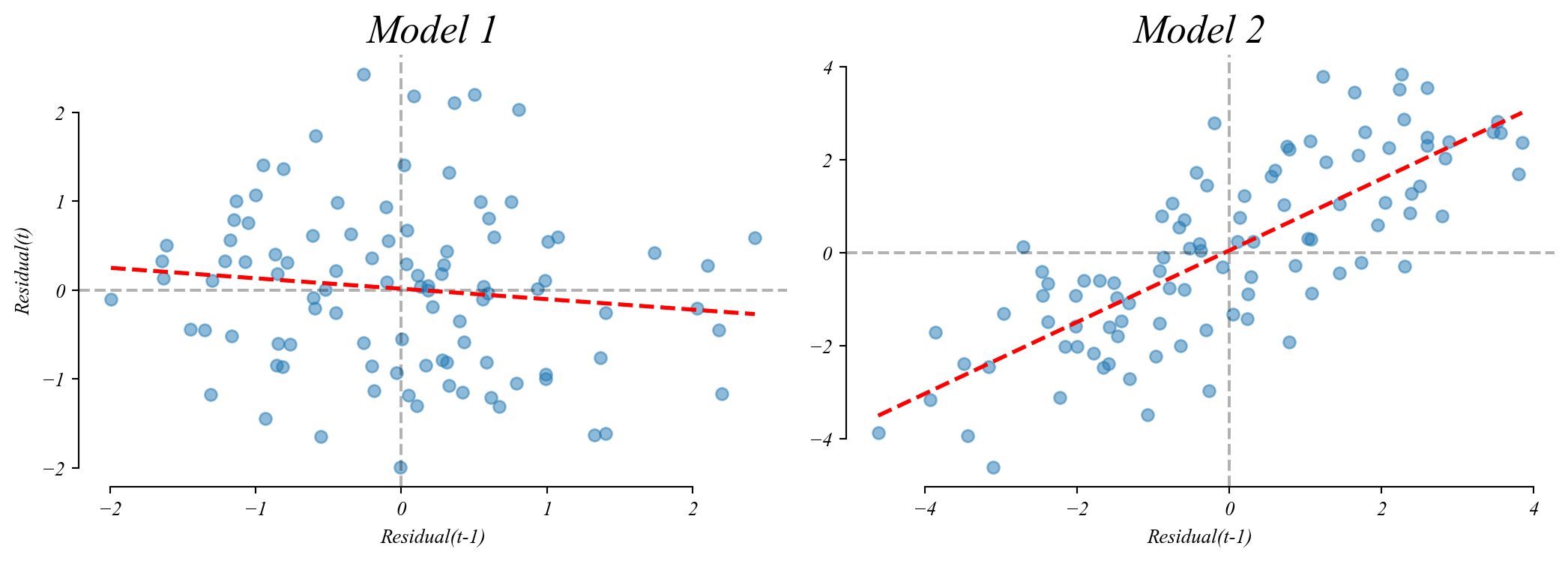
Model 1 has no (meaningful) relationship between consecutive residuals.
Model 2 shows a positive relationship (autocorrelation).
Checking Independence
Use a lagged residual plot to check for autocorrelation.
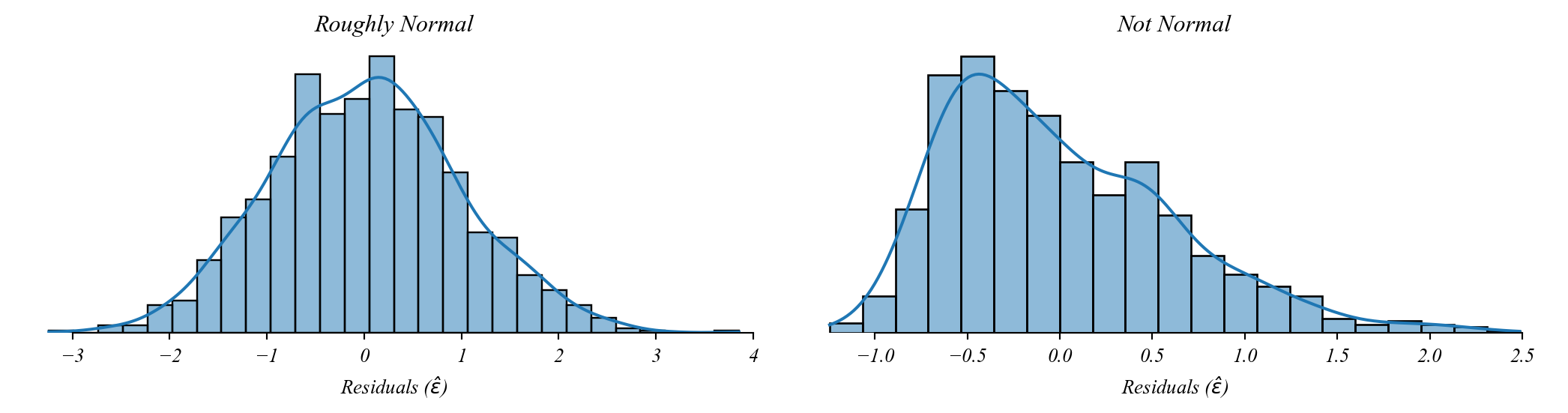
Assumption 4: Normality
Residuals should be normally distributed.
By the CLT we can still use GLM without this so long as the sample is large.
Looking Forward
Extending the GLM framework
Next Up:
- Part 4.4 | The Problem of Timeseries
Later:
- Part 5.1 | Numerical Controls
- Part 5.2 | Categorical Controls
- Part 5.3 | Interactions
- Part 5.4 | Model Selection
> all built on the same statistical foundation